Medicinal bacteria, also known as therapeutic bacteria or probiotics, are living microorganisms that provide health benefits when consumed or applied to the body. These bacteria can be found naturally in the human body, particularly in the digestive system, and also in certain foods and supplements.
Bacteria are intimately involved in our daily lives. In the past, these microbes have been employed to make cheese, yogurt, and wine. Microorganisms have been extensively exploited as microbial cell factories to produce polymers, feed for livestock, dietary supplements, and pharmaceuticals in more recent years through metabolic engineering.
However, pathogens like pneumonia, Salmonella, and Staphylococcus that cause a number of infectious diseases are also widely distributed together with these bacteria that are helpful to human health.
For the manufacture of high value-added compounds, it is crucial to be able to metabolically manage these helpful industrial microbes, and to modulate dangerous pathogens to decrease their pathogenic features.
A KAIST research team led by Distinguished Professor Sang Yup Lee of the Department of Biochemical Engineering had developed a new sRNA tool that can effectively inhibit target genes in various bacteria, including both gram-negative and gram-positive bacteria. The research results were published in Nature Communications.
In the past, we could only develop new tools for gene knockdown for each bacterium, but now we have developed a tool that works for a variety of bacteria.
Sang Yup Lee
Although sRNA is a powerful tool for synthesizing and controlling target genes in E. coli, it has been challenging to apply it to gram-positive bacteria that are useful for industry, such as Bacillus subtilis and Corynebacterium, in addition to gram-negative bacteria like E. coli..
The study team created a new sRNA platform that can successfully silence target genes in a variety of bacteria, including both gram-negative and gram-positive bacteria, to solve this issue. They surveyed thousands of microbial-derived sRNA systems in the microbial database, and eventually designated the sRNA system derived from Bacillus subtilis that showed the highest gene knockdown efficiency, and designated it as “Broad-Host-Range sRNA,” or BHR-sRNA.
A similar well-known system is the CRISPR interference (CRISPRi) system, which is a modified CRISPR system that knocks down gene expression by suppressing the gene transcription process.
The CRISPRi system’s Cas9 protein has a very large molecular weight, and there have been instances of bacterial growth suppression. The BHR-sRNA method created in this study demonstrated similar gene knockdown efficiency as CRISPRi while having no effect on bacterial growth.
The BHR-sRNA system was tested on 16 different gram-positive and gram-negative bacteria, and it was shown to be effective in 15 of them, demonstrating the BHR-sRNA system’s adaptability.
Additionally, it was shown that in 10 bacteria, the gene knockdown capability was superior to the current sRNA technique based on E. coli. The BHR-sRNA system has proven to be a versatile device that can successfully block gene expression in a variety of bacteria.
The BHR-sRNA was shown to inhibit the pathogenicity by suppressing the gene expressing the virulence factor in order to address the issue of antibiotic-resistant pathogens, which has recently become increasingly critical.
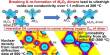
By using BHR-sRNA, biofilm formation, one of the factors resulting in antibiotic resistance, was inhibited by 73% in Staphylococcus epidermidis, a pathogen that can cause hospital-acquired infections. Antibiotic resistance was also weakened by 58% in the pneumonia-causing bacteria Klebsiella pneumoniae.
Additionally, BHR-sRNA was used to build microbial cell factories in order to produce high value-added compounds with improved production performance from industrial bacteria.
Notably, superior industrial strains were constructed with the aid of BHR-sRNA to produce the following chemicals: valerolactam, a raw material for polyamide polymers, methyl-anthranilate, a grape-flavor food additive, and indigoidine, a blue-toned natural dye.
This study’s creation of BHR-sRNA will hasten the commercialization of bioprocesses that create high value-added materials and chemicals including imitation meat, jet fuel, dietary supplements, medications, and polymers. It is also believed that it will aid in the eradication of germs that are resistant to antibiotics in order to get ready for a potential new pandemic.
“In the past, we could only develop new tools for gene knockdown for each bacterium, but now we have developed a tool that works for a variety of bacteria,” said Sang Yup Lee.