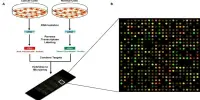
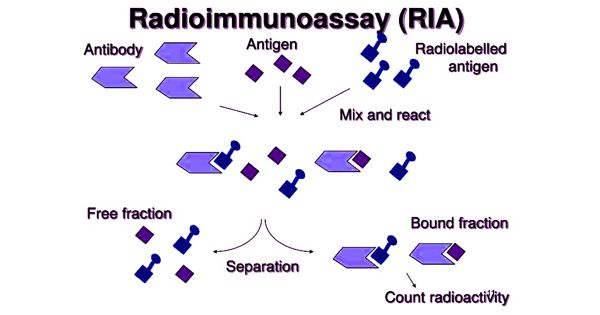
RNA sequencing, also known as RNA-seq, is a technique that allows researchers to study gene expression, alternative splicing, RNA editing, and other aspects of RNA biology. RNA-seq has a wide range of applications in biology and medicine, including the study of gene regulation, disease mechanisms, and drug discovery.
How do cells work in a normal state? How do they change when they cause disease? Do they react as desired to new drugs?
The specialized method known as single-cell RNA sequencing, or simply “scRNA-seq,” is now practically required for anyone looking for answers to these and other related problems in the laboratory.
This method gives a precise picture of gene expression in a single cell at a particular moment in time, together with the related regulatory networks, enabling inferences about the molecular underpinnings of cell activity.
A research team at the Julius-Maximilians-Universität Würzburg (JMU) has now further improved a single-cell RNA sequencing technique it previously developed for use in bacteria. This implies that laboratory work is completed even more quickly than before and with information that is far more accurate. The team presents its development in the journal mBio.
We were even able to detect sRNAs at the single cell level, which had not been possible before.
Professor Jörg Vogel
High throughput thanks to automation
The leader of the study now published is Professor Jörg Vogel. Vogel heads the Institute for Molecular Infection Biology (IMIB) at JMU and is director of the Helmholtz Institute for RNA-based Infection Research (HIRI). He is one of the world’s leading experts in the field of RNA research.
“By integrating a user-friendly and highly flexible automation process, we have achieved higher cell throughput,” says Vogel, describing one advantage of the method now being presented.
The method also works more reliably, lowering the failure rate when attempting to read genetic data, and offers more details on the gene expression of specific cells at lower sequencing costs.
Averages hide important details
Until a few years ago, studies of the transcriptome the set of all genes active in a cell at a given time in bacteria relied on bulk RNA sequencing (RNA-seq).
“However, this approach only provides average values of a cell population and therefore does not allow any conclusions to be drawn about possible differences between individual bacteria within this population,” Vogel explains.
But these variations which scientists in this instance refer to as “phenotypic heterogeneity” are frequently observed in bacteria. They play a crucial part in bacterial survival strategies as they allow them to quickly adapt to shifting conditions.
Bacteria pose special challenges for technology
For eukaryotic cells with a nucleus, single-cell RNA sequencing was first made available in 2009; however, the development of this method for bacterial cells has progressed far more slowly. A number of challenges are responsible for this: “Prokaryotic cells are much smaller compared to eukaryotes, which means you have much less material to study per cell,” Vogel explains.
Cell lysis, the process of destroying the cell wall, and identifying certain bacterial transcripts are further issues.
It is true that recent technological developments have also made bacterial single-cell transcriptomics a reality. Nevertheless, there is room for improvement, for example, because the frequency of cell loss is too high or short transcripts, such as regulatory small RNAs (sRNAs), are poorly detected or cannot be measured at all.
“In addition, transcript recognition is currently limited to about 200 genes per cell, which is far below the average bacterial transcriptome,” Vogel explains.
Successful validation on Salmonella
The research team was able to demonstrate that some of these issues can be resolved by the new advancements in scRNA sequencing technology by examining Salmonella-type bacteria under various growth circumstances. The statistics demonstrate that the applied adjustments reduced cell loss while increasing cell throughput and the protocol’s robustness.
In addition, the scientists were able to improve gene coverage and gene detection limits.
“We were even able to detect sRNAs at the single cell level, which had not been possible before,” Vogel says. This, he adds, will allow exploration of the regulatory functions of sRNA at the single-cell level in future studies.
The data also support the heterogeneity within a single cell population, which could not be inferred from the average values of earlier sequencing methods. They now offer details, for instance, on the activity of genes that are crucial for the ability of these bacteria to cause disease.
As a result, the technique is particularly well suited for investigations with little beginning material, such as those that analyze discrete populations of intracellular bacteria or small subpopulations of bacterial cells in host niches.