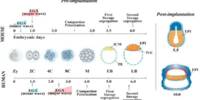
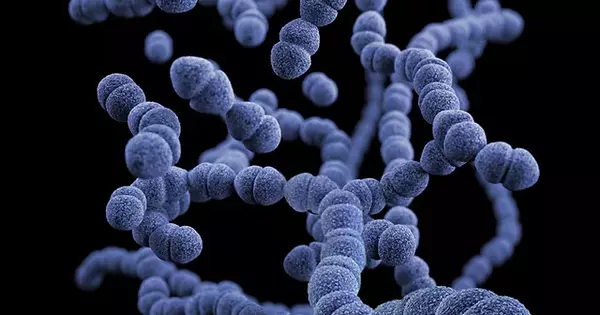
Extensive research on a recalcitrant bacterium could result in better probiotics. New research from North Carolina State University shows progress in gathering data on an important – but difficult to characterize – human gut bacterium called Bifidobacterium, which is used in many probiotics that help maintain healthy microbiomes. The findings have the potential to improve so-called “good bacteria.”
“As our lab expands and diversifies the types of good bacteria we work with, we turn to more finicky bacteria, like Bifidobacterium,” said Rodolphe Barrangou, the Todd R. Klaenhammer Distinguished Professor of Food, Bioprocessing, and Nutrition Sciences at NC State and lead author of a paper describing the research published today in Proceedings of the National Academy of Sciences.
“While this bacterium is more difficult to grow and work with than others, we were able to make some important discoveries and learn more about the bacterium’s genetic basis for its health-promoting functions.”
Bifidobacterium is one of the two main players in the probiotic industry, along with Lactobacillus, and is particularly dominant in the colon of infants. However, it is much more difficult to manipulate than Lactobacillus.
Meichen (Echo) Pan
“Bifidobacterium is one of the two main players in the probiotic industry, along with Lactobacillus, and is particularly dominant in the colon of infants,” said Meichen (Echo) Pan, an NC State Ph.D. student and the paper’s first author. “However, it is much more difficult to manipulate than Lactobacillus.”
NC State researchers discovered the bacterium’s internal CRISPR-Cas system as well as a portable engineered CRISPR effector. CRISPR-Cas systems are adaptive immune systems that enable bacteria to withstand attacks from enemies such as viruses. Scientists have adapted these systems to remove or cut and replace specific genetic code sequences.
It turns out that Bifidobacterium has a plethora of native CRISPR-Cas systems, one of which is a relatively unknown type I-G system. In separate experiments, the researchers used this internal system and a portable Cas effector called a cytosine base editor to resensitize a Bifidobacterium strain to a common antibiotic called tetracycline. Many bacteria are naturally resistant to antibiotics.
“Restoring antibiotic sensitivity is conceptually and practically important because bacteria can potentially transfer antibacterial resistance to other bacteria in the gut,” Pan said.
The researchers also discovered tiny changes in different strains of the bacterium, known as single nucleotide polymorphisms or SNPs, that appeared to reflect large differences in the strains’ phenotypes, or characteristics.
“This was an unexpected lesson: One letter difference in strains with genetic codes that are over 99 percent similar can make huge differences,” said Barrangou. “What genes turn on and how they behave as a result of their environment can make a significant difference, and researchers will need to customize the CRISPR tool to adapt the editing strategy accordingly.”
Scientists believe that our increasingly sterile environment is eroding this microbial defense. Our homes are cleaner, and we come into less contact with bacteria-infested soil and animals than our forefathers did. Our food and water are also treated in ways that keep microscopic parasites at bay. These advancements have undoubtedly reduced our exposure to potentially harmful microorganisms, but they may also be limiting the microbial diversity that aids in the maintenance of our immune systems.
But perhaps the most serious threat to our microbiome is our overuse of antibiotics. These drugs are highly valued for their ability to treat bacterial infections. Unfortunately, they frequently kill beneficial bacteria alongside harmful bacteria.