Transcriptomics is defined as the study of the transcriptome—the entire set of RNA. It is also known as expression profiling because it is a study of the expression levels of mRNAs in a given cell population. The decision to extend or terminate the RNA chain at specific DNA template positions during transcription is kinetically regulated, but the methods used to measure the rates of these processes are not quantitative enough to allow a detailed mechanistic analysis of the steps involved.
The more we understand cellular processes like RNA regulation, the better molecular therapies we can develop. Until now, tracking the regulation of non-coding RNA, or RNA that is not further converted into proteins, has been particularly difficult. A team of researchers from Helmholtz Munich and the Technical University of Munich (TUM) has developed a minimally invasive reporter system that allows for highly sensitive monitoring of both coding and non-coding RNA production.
For cellular processes, our genetic DNA information is transcribed into RNA, which is then processed further before serving as a blueprint for proteins or performing a cellular function on its own. The types of RNA produced and the quantities produced reveal a lot about the health of our cells. Cells, for example, produce more RNA molecules that code for proteins involved in the immune response during an infection.
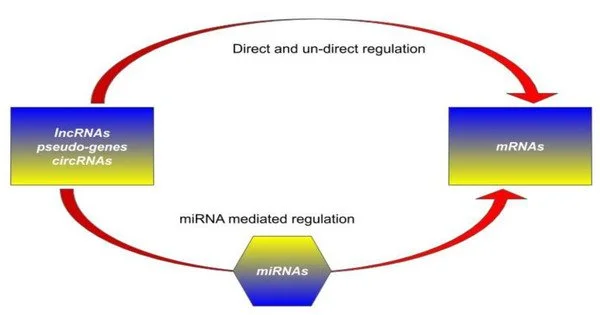
Researchers can use existing reporter systems to monitor the process of DNA molecules being translated into proteins via RNA. Not all human genes, however, code for proteins. The vast majority of human genes are non-coding, including long non-coding RNA genes (lncRNA). These are RNA molecules made up of more than 200 building blocks that do not serve as protein blueprints. Instead, they regulate critical cell processes. According to preliminary research, lncRNA is involved in processes such as regulating RNA production, organizing structures in the cell nucleus, and switching specific enzymes on and off.
Unlike previous methods, INSPECT encodes sequences for reporter proteins in modified introns. These are sequences in the pre-mature RNA molecule that are removed naturally and eliminated by the cell during processing. INSPECT stabilizes the introns such that, rather than being degraded after removal, they are transported to the cellular cytoplasm where they are translated into reporter proteins.
Dong-Jiunn Jeffery Truong
Despite their importance for cellular processes, it has been difficult to investigate lncRNAs with existing methods. So far, this was only partially possible, for example in fixed cells at specific time points, because classical reporter systems based on the translation into proteins cannot be used.
INSPECT permits the monitoring of non-coding RNA
A solution has now been found in the form of a new reporter system: INSPECT. A team working with Gil Westmeyer, Professor of Neurobiological Engineering at TUM and the Director of the Institute for Synthetic Biomedicine at Helmholtz Munich, has now published the newly developed reporter system in the journal Nature Cell Biology.
“Unlike previous methods, INSPECT encodes sequences for reporter proteins in modified introns. These are sequences in the pre-mature RNA molecule that are removed naturally and eliminated by the cell during processing. INSPECT stabilizes the introns such that, rather than being degraded after removal, they are transported to the cellular cytoplasm where they are translated into reporter proteins,” explains first author Dong-Jiunn Jeffery Truong. The researchers can then use conventional methods to detect reporter protein signals such as fluorescence.
INSPECT modifies neither the completed RNA nor the proteins
Thus, the new molecular biology tool not only solves the problem of tracking the generation of non-coding RNA, but also provides advantages for studying coding RNA. Current reporter systems frequently risk damaging the RNA or proteins under investigation, for example, because they must be fused directly to the RNA being studied in order to be co-translated into proteins. INSPECT modifies introns rather than the completed RNA or proteins.
The team demonstrated the function of INSPECT using various examples of coding and non-coding RNA. They monitored the production of RNA for interleukin 2, a protein that is produced in greater quantities in response to infections. They have also achieved highly sensitive monitoring of the production of two lncRNAs and tracked changes in regulation during the investigation period.
“INSPECT adds an important molecular biology tool to the biomedical toolbox, making it easier to study the role of certain non-coding RNA molecules in cell development and to investigate how their regulation can be modulated, for example, to prevent them from turning into cancer cells,” says Prof. Westmeyer. “With the minimally invasive reporter system EXSISERS, which we previously developed to study protein isoforms, it may be possible in the future to study an entire genetic regulation process in living cells, from RNA processing to the production of specific protein variants.”