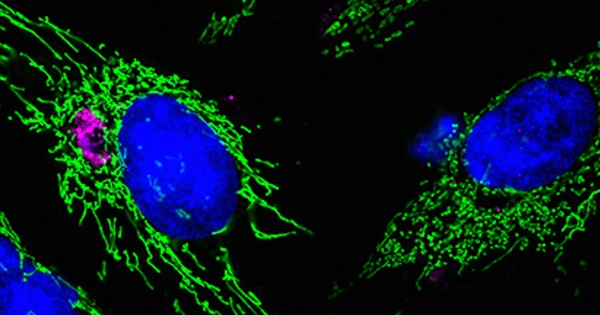
Chlamydiae are a group of obligate intracellular bacteria that have a unique life cycle, characterized by both a replicative, reticulate stage inside host cells and a transmissive stage outside of host cells. This life cycle has led to their classification as a separate phylum, called Chlamydiae, which is distinct from both Gram-negative and Gram-positive bacteria.
One of the notable features of Chlamydiae is that they have evolved to live and replicate within host cells, and they have lost many of the genes that other bacteria use to survive outside of host cells. This has led scientists to believe that Chlamydiae have a specialized adaptation for intracellular life, and provides an interesting view on how bacteria can evolve to live within host cells. Additionally, their unique biology, which involves a unique developmental cycle and a lack of metabolic pathways, makes them a model system to study the intracellular lifestyle and to understand the mechanisms of host-pathogen interaction.
Today, all chlamydiae live within the cells of hosts ranging from amoeba to animals. A team of researchers from the University of Vienna and Wageningen University & Research discovered that chlamydiae’s ancestor most likely already lived inside host cells, but that chlamydiae infecting amoeba evolved later in unexpected ways for intracellular bacteria. The study, which was published in the journal Nature Microbiology, is an important step toward understanding the origins and evolution of endosymbiotic bacteria, including human pathogens.
We’ve only recently gained the capability to sequence genomes directly from environmental samples to explore the breadth of chlamydial diversity,
Matthias Horn
Chlamydiae are best known for the human pathogen Chlamydia trachomatis, but this bacterial family evolved over a billion years ago, long before the first animals appeared. Nonetheless, all chlamydiae discovered today live inside a wide variety of hosts, ranging from small amoeba to animal cells. But, since the first chlamydiae genomes were sequenced 20 years ago, scientists have been puzzled by the fact that, while chlamydiae that infect animals have small genomes similar to other endosymbionts, those that infect amoeba have larger genome sizes more similar to free-living bacteria. The inability to grow these microbes in the lab has hampered research into the evolution of this diverse bacterial group.
The teams around Matthias Horn (University of Vienna) and Thijs Ettema (Wageningen University & Research) could circumvent this problem: “We’ve only recently gained the capability to sequence genomes directly from environmental samples to explore the breadth of chlamydial diversity,” explain the researchers. With this new data in hand, they then traced back the evolution of chlamydiae.
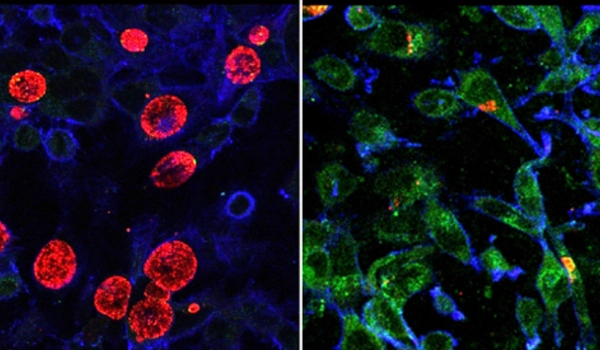
Using state-of-the-art computational methods, they reconstructed the genome of the last common ancestor of all known chlamydiae. The researchers found that “this extinct microbe had all the genes needed to be an endosymbiont. Even genes important for chlamydial animal pathogens today were likely already present.” This means that chlamydiae have been infecting host cells for over a billion years of evolutionary history.
However, to their surprise, the research team also found that chlamydiae infecting amoeba gained many metabolic genes only later, despite the fact that endosymbionts have fewer opportunities to exchange genes with other bacteria. “Our results show that more gene exchange happened in some chlamydiae than expected for endosymbionts,” the authors explain, “including the gain of key metabolic genes.”
This result challenges how we think about the evolution of endosymbionts. But the researchers also suggest a solution to this conundrum: “It’s not so surprising when you think about the environment these chlamydiae live in: Amoeba often host multiple endosymbionts and feed on free-living bacteria, so there are other microbes around increasing the accessible gene pool. In addition, most chlamydiae move between different hosts, and exposure to changing environments could explain why it might be beneficial for these endosymbionts to keep and even gain additional metabolic genes.”
Scientists are curious to see whether this mode of endosymbiont evolution is more widespread. In any case, this study is an important step for understanding the emergence and evolution of endosymbiotic bacteria, including human pathogens.